Contents
When a user used a partical map region ("boxed map") by phenix.map_box or relion_image_handler, cryo_fit's automatic mrc to sit map format conversion may not work properly.
Therefore, please use situs map2map to convert your mrc format map to situs format map. You can convert by "map2map user.mrc user.sit" then enter 1 for "Convert to classic Situs (auto)*". Then, provide this user.sit file to your cryo_fit. For example, phenix.cryo_fit user.pdb user.sit
When Doo Nam provided situs made sit map file, the cryo_fit ran smoothly again.
When a user didn't use phenix.map_box, it means that the map dimensions need to be larger.
Like other MD simulations, gromacs need enough map box size to cover the atomic model to run (ziggle and wiggle). Refer Waters seems to be out of the box
For example, stuck-out red oxygen atoms outside the right edge of the box are the problem.
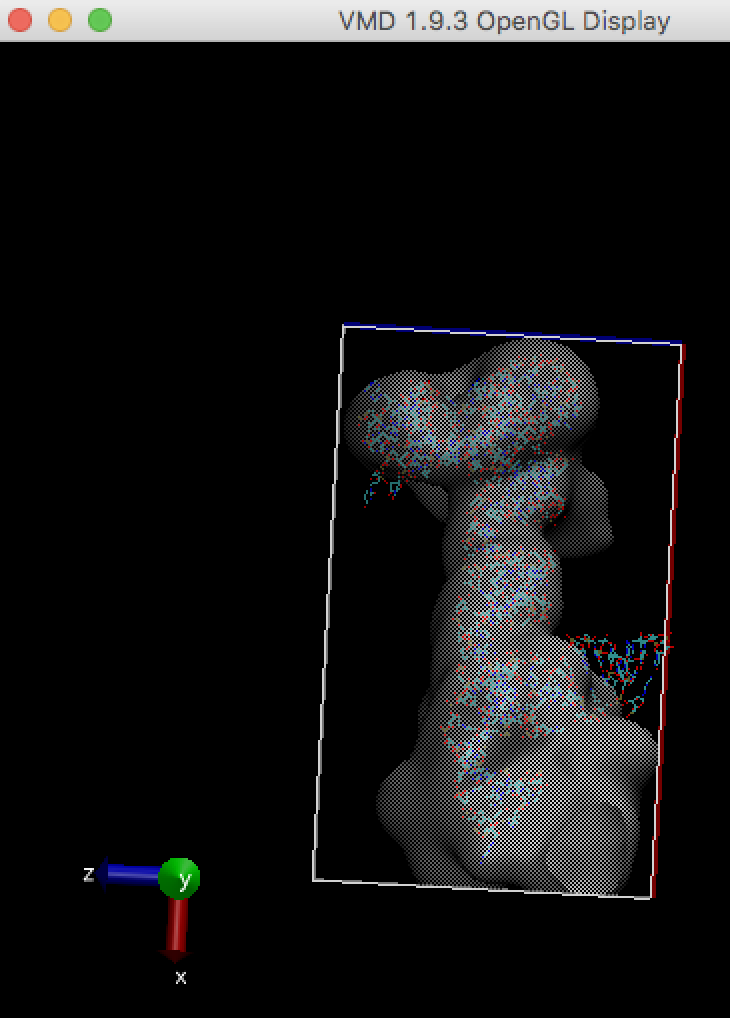
In order to run any MD simulation (including cryo_fit), a box should be large enough like
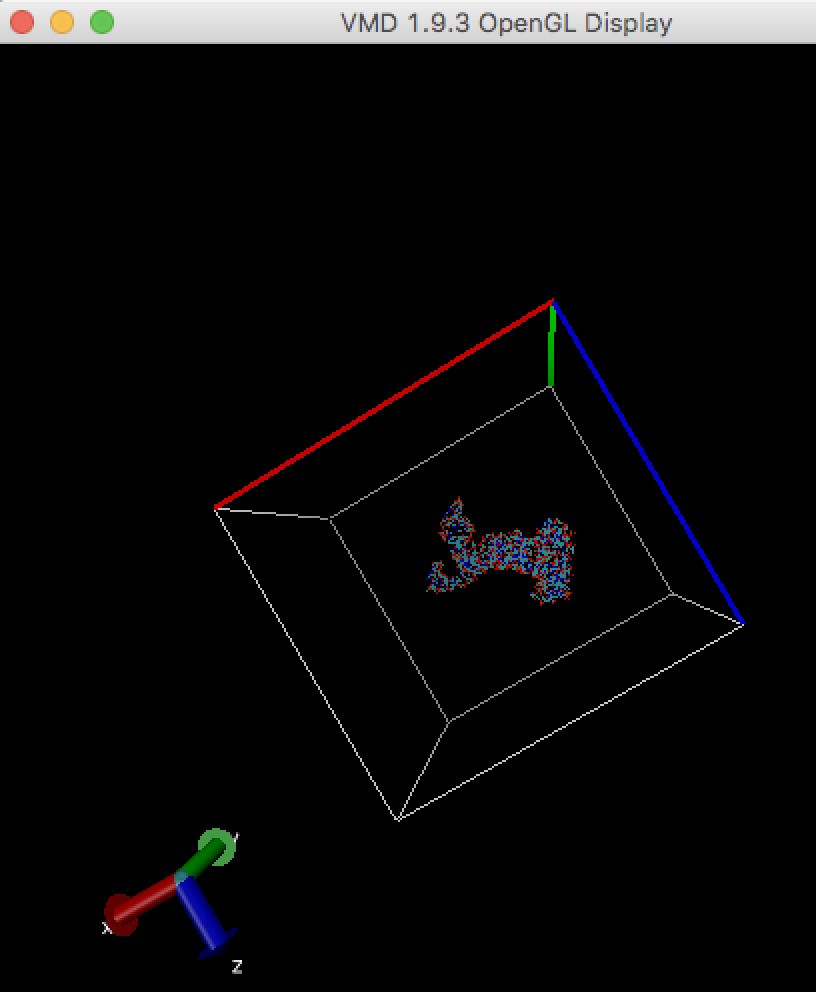
Make map box size larger (see "How to enlarge map box size?" above), and run cryo_fit again. You can check map box size by VMD. Alternatively, remove sticking out atoms if these are unnecessary, then run cryo_fit again.
For protein modeling, I would use cryo_fit2 which is not limited by box size requirement. It better fits than cryo_fit1 in terms of fitting and geometry anyway.